PCA & Low-Rank Approximation
From variance maximization to modern dimensionality reduction — the mathematical theory behind the most widely used unsupervised learning method
Overview & Motivation
The SVD established that the right singular vectors of a centered data matrix are the principal components, and that the Eckart–Young–Mirsky theorem makes the truncated SVD the optimal low-rank approximation. This topic develops PCA as a standalone framework — starting from the variance-maximization problem, connecting it to the SVD and the Spectral Theorem, and extending it to the modern variants that handle nonlinear, corrupted, and high-dimensional data.
Principal Component Analysis (PCA) finds the orthogonal directions of maximum variance in a dataset (rows = observations, columns = features). It answers a simple question: if we could keep only directions, which directions capture the most information?
The answer is the top eigenvectors of the sample covariance matrix — equivalently, the top right singular vectors of . This topic covers:
- Variance maximization: PCA as a constrained optimization problem (Rayleigh quotient).
- Optimal projection: the Eckart–Young connection — PCA is the best -dimensional linear projection.
- The scree plot: heuristics for choosing (elbow, Kaiser, parallel analysis).
- Kernel PCA: extending PCA to nonlinear manifolds via the kernel trick.
- Probabilistic PCA: a latent variable model that handles missing data and provides a generative framework.
- Robust PCA: separating low-rank structure from sparse corruption (Principal Component Pursuit).
- Sparse PCA: interpretable principal components via -penalized loadings.
What We Cover
- PCA from First Principles — the Rayleigh quotient and sequential variance maximization.
- PCA via the SVD — the correspondence table and numerical superiority.
- Geometric Interpretation — optimal projection and the dual view (variance = error minimization).
- The Scree Plot — dimensionality selection (elbow, Kaiser, parallel analysis).
- Low-Rank Approximation — the Eckart–Young connection and effective rank.
- Computing PCA in Practice — scikit-learn classes and the standardization question.
- The Biplot — visualizing observations and variables together.
- Kernel PCA — nonlinear dimensionality reduction via the kernel trick.
- Probabilistic PCA — the latent variable model and EM for missing data.
- Robust PCA — Principal Component Pursuit for low-rank + sparse decomposition.
- Sparse PCA — interpretable loadings via regularization.
PCA from First Principles: Variance Maximization
The Setup
Let be a centered data matrix ( observations, features, column means subtracted). The sample covariance matrix is:
This matrix is symmetric and positive semi-definite. By the Spectral Theorem, it has an orthogonal eigendecomposition:
with eigenvalues and orthonormal eigenvectors .
The Variance-Maximization Problem
The first principal component is the unit vector that maximizes the variance of the projection :
This is a Rayleigh quotient problem. By the Spectral Theorem (specifically, the Courant–Fischer minimax principle), the maximum is achieved at (the top eigenvector), and the maximum variance is .
Proposition 1 (Sequential variance maximization).
The -th principal component is the unit vector that maximizes subject to and . The solution is (the -th eigenvector of ), and the maximum variance is .
Proof.
By induction on the Spectral Theorem’s eigenvalue ordering.
After deflation, we project onto the orthogonal complement of . Any vector in this subspace can be written as with . Then:
with equality when and for , i.e., . This is the restricted Rayleigh quotient, and the bound follows from the Spectral Theorem’s eigenvalue ordering .
∎The Principal Components
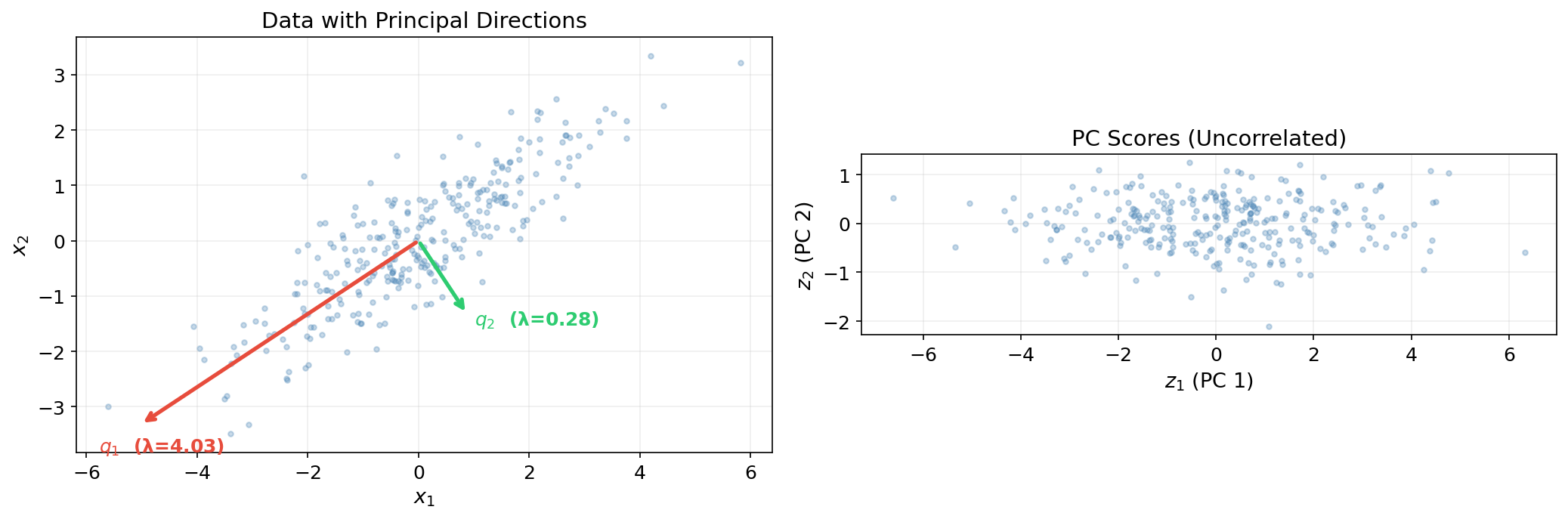
The principal components (or PC scores) are the projections of the data onto the eigenvectors:
where is the -th observation’s coordinate along the -th principal direction. The columns of are uncorrelated with variances :
This diagonalization — transforming correlated features into uncorrelated components ordered by variance — is the essence of PCA.
PCA via the SVD
The SVD Connection
The SVD topic §8 established the fundamental connection. We now develop it fully.
Let be the SVD of the centered data matrix. Then:
Comparing with the Spectral Theorem decomposition :
Theorem 1 (PCA via SVD).
Let be a centered data matrix with SVD . The principal components are the columns of , the PC scores are the columns of , and the -th eigenvalue of the sample covariance matrix is .
| PCA quantity | Spectral Theorem | SVD |
|---|---|---|
| Principal directions | Eigenvectors of | Right singular vectors of |
| Eigenvalues (variances) | ||
| PC scores | ||
| Explained variance ratio |
Numerical Superiority of the SVD
Forming explicitly squares the condition number: . For ill-conditioned data, the smallest eigenvalues can be lost to floating-point roundoff. The SVD computes the same decomposition directly on , preserving numerical accuracy. This is why every serious PCA implementation uses the SVD internally.
Definition 1 (Condition number for PCA).
The condition number of PCA is (the ratio of the largest to smallest singular value of ). When is large, the data is ill-conditioned for PCA — the smallest principal components are unreliable.
import numpy as np
# PCA via eigendecomposition vs. SVD: verify the connection
np.random.seed(42)
n, d = 500, 3
cov_true = np.array([[3.0, 1.5, 0.5], [1.5, 2.0, 0.3], [0.5, 0.3, 0.5]])
X = np.random.multivariate_normal(np.zeros(d), cov_true, n)
X_centered = X - X.mean(axis=0)
# Method 1: Spectral Theorem on covariance
Sigma_hat = (X_centered.T @ X_centered) / (n - 1)
evals_eig, Q = np.linalg.eigh(Sigma_hat)
idx = np.argsort(evals_eig)[::-1]
evals_eig, Q = evals_eig[idx], Q[:, idx]
# Method 2: SVD of centered data
U, sigma, Vt = np.linalg.svd(X_centered, full_matrices=False)
evals_svd = sigma**2 / (n - 1)
# Eigenvalues match: λ_i = σ_i² / (n-1)
for i in range(d):
print(f" λ_{i+1}: eigendecomp = {evals_eig[i]:.6f} SVD = {evals_svd[i]:.6f}")
# Condition numbers: SVD avoids squaring
print(f"\n κ(X) = {sigma[0]/sigma[-1]:.2f}")
print(f" κ(X^T X) = {evals_eig[0]/evals_eig[-1]:.2f} ≈ κ(X)² = {(sigma[0]/sigma[-1])**2:.2f}")Geometric Interpretation
PCA as Optimal Linear Projection
PCA finds the -dimensional subspace that best preserves the data’s variance. Equivalently, it finds the projection that minimizes reconstruction error.
Theorem 2 (PCA as optimal projection).
Let be centered data with SVD . The -dimensional affine subspace that minimizes the total squared reconstruction error is spanned by the top right singular vectors :
This is the Eckart–Young–Mirsky theorem applied to the centered data matrix.
Proof.
The reconstruction error for a rank- projection is . We need to show this is minimized when .
Expand using the SVD :
where the last equality uses the unitary invariance of the Frobenius norm ( for orthogonal ).
Since projects onto the orthogonal complement of the -dimensional subspace, the minimum is achieved when projects onto the span of the top right singular vectors. The error equals — the sum of the squared singular values we discard. This is exactly the Eckart–Young theorem.
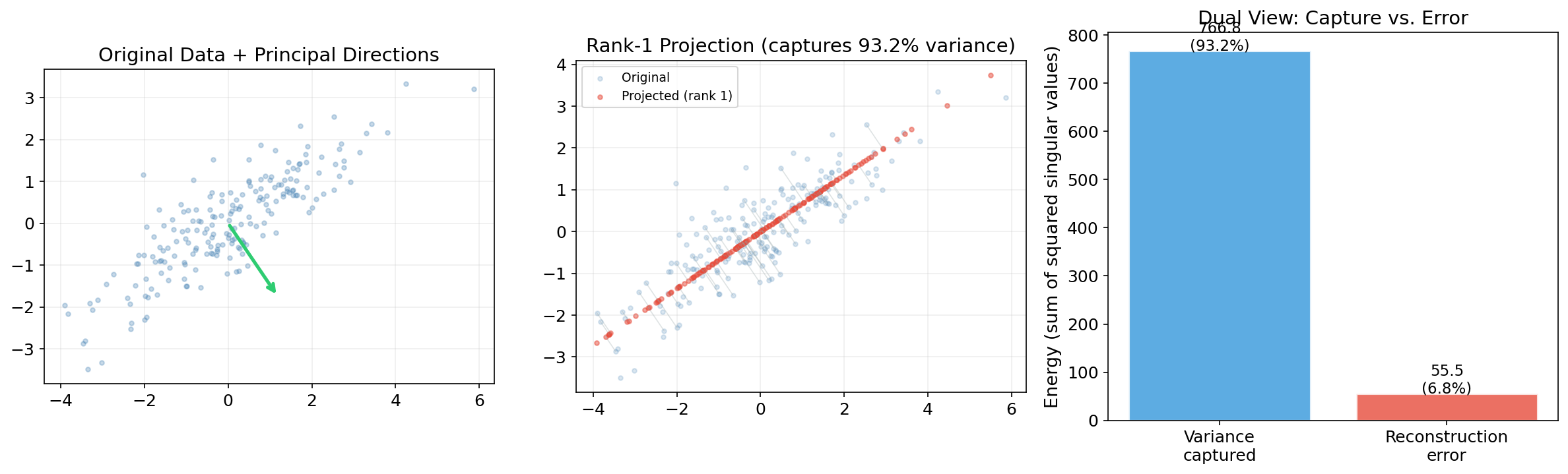
∎The Dual View: Variance Maximization = Error Minimization
PCA simultaneously:
- Maximizes the variance of the projected data: .
- Minimizes the reconstruction error: .
Since total variance is constant, maximizing (1) is equivalent to minimizing (2). This dual view is central to understanding dimensionality reduction.
Theorem 3 (Optimal projection subspace).
Among all -dimensional subspaces , the one that minimizes the total squared distance from the data to its projections is the span of the top principal components:
where is the orthogonal projection onto .
The Reconstruction Formula
Given the top principal components and PC scores , the rank- reconstruction is:
The projection matrix is the orthogonal projector onto the principal subspace.
The Scree Plot and Dimensionality Selection
The Scree Plot
The scree plot displays the eigenvalues (or singular values, or explained variance ratios) in descending order. The goal is to identify the “elbow” — the point where the eigenvalues transition from large (signal) to small (noise).
Definition 2 (Explained variance ratio).
The explained variance ratio for components is:
where are eigenvalues of the covariance matrix and are singular values of the centered data matrix.
Selection Criteria
Several heuristics exist for choosing the number of components :
| Method | Rule | When to use |
|---|---|---|
| Elbow method | Choose at the “bend” in the scree plot | Visual; subjective but widely used |
| Kaiser criterion | Keep components with (for standardized data) | Only for correlation-matrix PCA |
| Explained variance threshold | Choose smallest s.t. (e.g., ) | When a target variance fraction is needed |
| Parallel analysis | Compare eigenvalues to those from random data of the same dimensions | Most statistically principled |
Parallel Analysis
Parallel analysis (Horn, 1965) generates many random datasets of the same size () from i.i.d. standard normals, computes their eigenvalues, and retains only those components whose observed eigenvalues exceed the 95th percentile of the random eigenvalues. This controls for the “eigenvalue inflation” that occurs in finite samples.
Eigenvalue spectrum
Cumulative explained variance
Selected k = 2 — explains 55.4% of total variance.(Click a bar to change k.)
Criteria comparison: Kaiser = 3 | Parallel analysis = 2 | Elbow = 2 | 95% threshold = 10
from sklearn.datasets import load_wine
from sklearn.preprocessing import StandardScaler
# Load and standardize the Wine dataset (13 features)
wine = load_wine()
X_wine = StandardScaler().fit_transform(wine.data)
n, d = X_wine.shape
# PCA via SVD
U, sigma, Vt = np.linalg.svd(X_wine, full_matrices=False)
eigenvalues = sigma**2 / (n - 1)
evr = eigenvalues / eigenvalues.sum()
cumulative_evr = np.cumsum(evr)
# Parallel analysis: compare to eigenvalues from random data
n_permutations = 100
random_eigenvalues = np.zeros((n_permutations, d))
for j in range(n_permutations):
X_random = np.random.randn(n, d)
_, s_rand, _ = np.linalg.svd(X_random, full_matrices=False)
random_eigenvalues[j] = s_rand**2 / (n - 1)
random_95th = np.percentile(random_eigenvalues, 95, axis=0)
k_parallel = np.sum(eigenvalues > random_95th)
# Selection results
k_kaiser = np.sum(eigenvalues > 1.0)
k_95 = np.searchsorted(cumulative_evr, 0.95) + 1
print(f"Kaiser criterion (λ > 1): k = {k_kaiser}")
print(f"95% variance threshold: k = {k_95}")
print(f"Parallel analysis: k = {k_parallel}")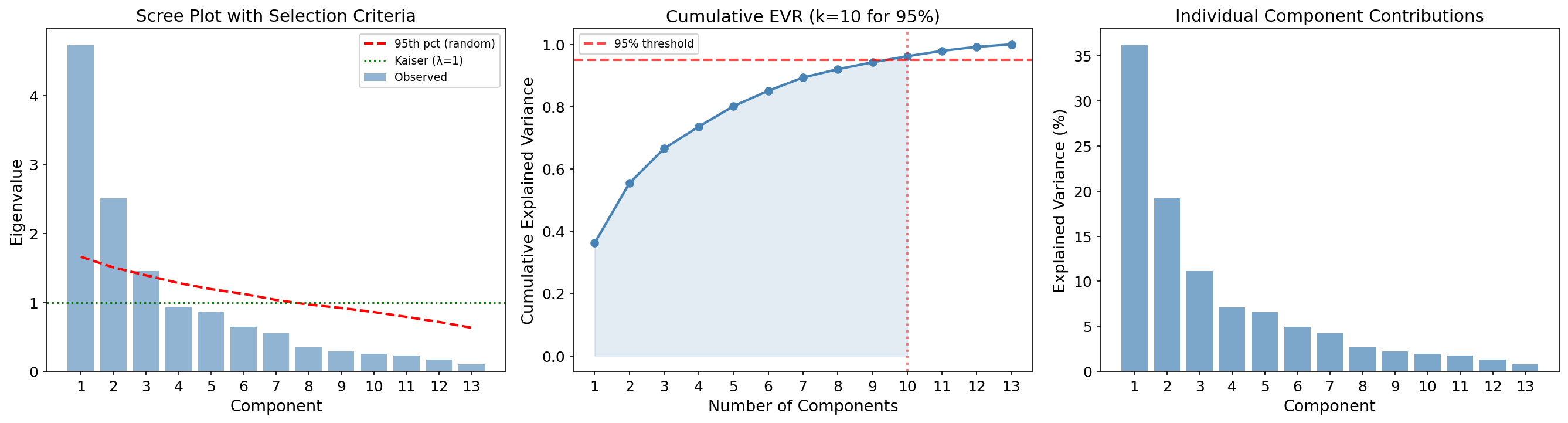
Low-Rank Approximation as Optimal Projection
The Eckart–Young Connection
The Eckart–Young–Mirsky theorem states that the best rank- approximation to is the truncated SVD . Applied to the centered data matrix :
This is exactly the PCA reconstruction: project onto the top PCs, then reconstruct. The Eckart–Young theorem guarantees this is optimal among all rank- approximations.
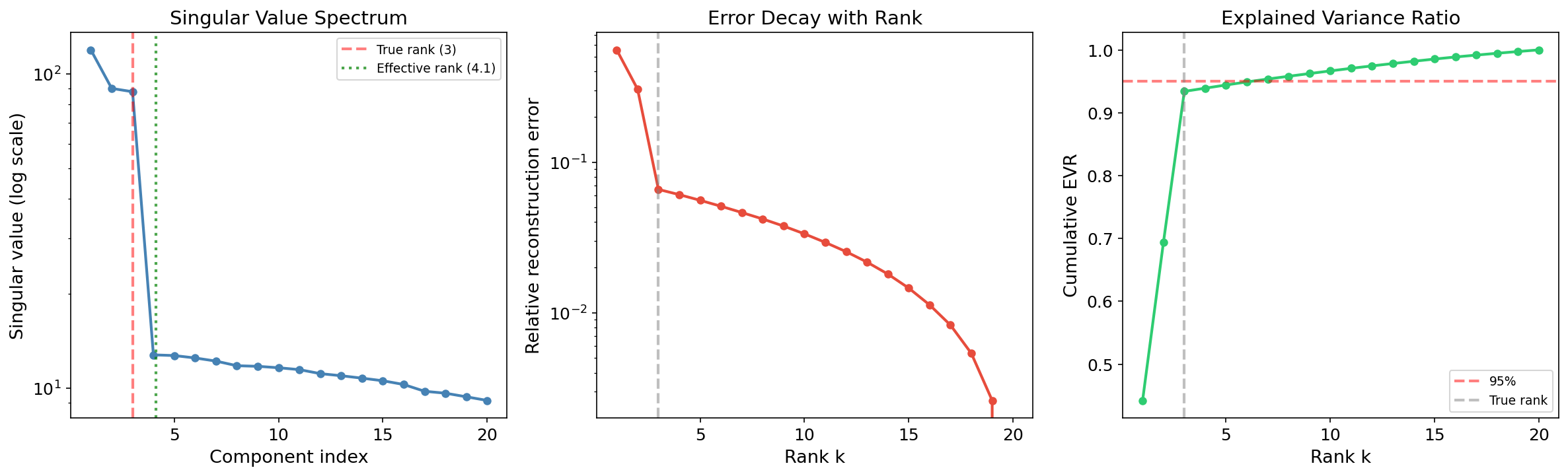
Effective Rank
Definition 3 (Effective rank).
The effective rank of a matrix measures how many singular values contribute meaningfully. Two common definitions:
- Hard threshold: for some tolerance .
- Entropy-based (Roy & Vetterli, 2007): where .
When the effective rank is much smaller than , the data is approximately low-rank and PCA is highly effective.
Remark.
The entropy-based effective rank has a nice information-theoretic interpretation: it measures the “uniformity” of the singular value distribution. When all singular values are equal, (full rank). When one singular value dominates, .
Computing PCA in Practice
scikit-learn
| Class | When to use | Algorithm |
|---|---|---|
PCA(n_components=k) | Moderate data; exact PCA | Full SVD via LAPACK or randomized SVD (auto) |
PCA(svd_solver='randomized') | Large , small | Halko–Martinsson–Tropp randomized SVD |
IncrementalPCA(n_components=k) | Out-of-core / streaming | Incremental SVD update |
TruncatedSVD(n_components=k) | Sparse matrices (text, graphs) | ARPACK or randomized |
Remark.
PCA centers the data automatically. TruncatedSVD does not center — use it for sparse data where centering would destroy sparsity (e.g., TF-IDF matrices in LSA).
Standardize or Not?
- Standardize (scale to unit variance) when features have different units or very different scales. PCA on the correlation matrix where .
- Don’t standardize when features are commensurable (same units, similar scales) — PCA on the covariance matrix preserves the natural scaling.
The Kaiser criterion () only makes sense when PCA is computed on the correlation matrix (standardized data), because the average eigenvalue of a correlation matrix is exactly 1.
Application: High-Dimensional Data Visualization
The Biplot
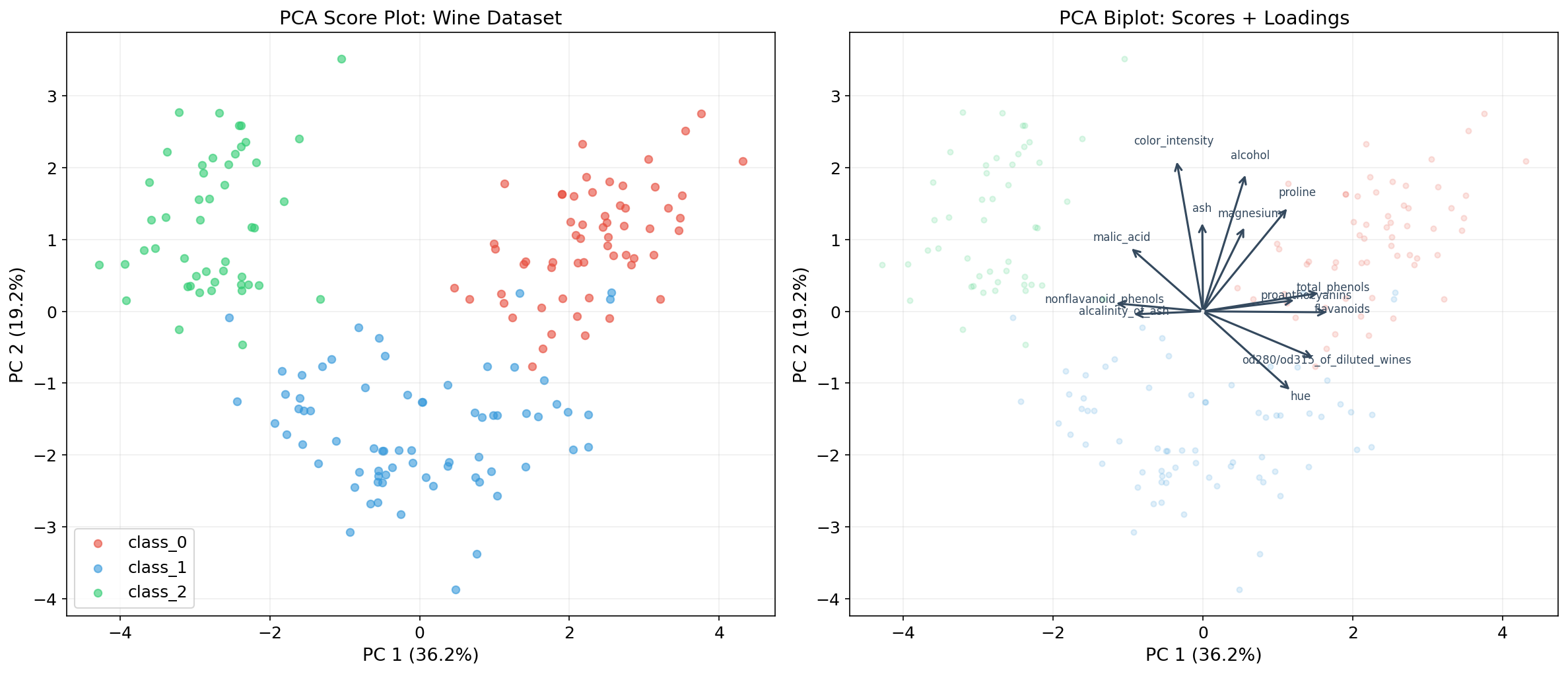
The biplot (Gabriel, 1971) simultaneously displays observations (PC scores) and variables (loadings) in the principal component space. It is the standard visualization for PCA results:
- Observations are plotted as points using PC scores .
- Variables are plotted as arrows (loading vectors) showing each original feature’s contribution to the PCs.
- Arrow length indicates the feature’s variance explained by the displayed PCs.
- Arrow angle between two features approximates their correlation (small angle ≈ high positive correlation; 90° ≈ uncorrelated; 180° ≈ high negative correlation).
Interpreting Loadings
The loadings are the entries of the principal component vectors . The loading tells us how much feature contributes to PC . Large loadings (in absolute value) indicate features that are important for that component.
Kernel PCA
The Limitation of Linear PCA
Linear PCA finds directions of maximum variance in the original feature space. But many real datasets have nonlinear structure — data lying on a curved manifold (Swiss roll), concentric circles, or a spiral. Linear PCA cannot “unfold” these structures.
The Kernel Trick
Kernel PCA (Schölkopf, Smola & Müller, 1998) applies PCA in a high-dimensional (possibly infinite-dimensional) feature space without ever computing explicitly:
- Compute the kernel matrix where is a kernel function.
- Center the kernel matrix: .
- Eigendecompose: (Spectral Theorem, since is symmetric PSD).
- The kernel PC scores are for the top eigenvectors.
Definition 4 (Common kernels for Kernel PCA).
| Kernel | Formula | Effect |
|---|---|---|
| Linear | Standard PCA | |
| RBF (Gaussian) | Smooth nonlinear embedding | |
| Polynomial | Polynomial feature interactions |
The RBF kernel with bandwidth is the most commonly used; it implicitly maps to an infinite-dimensional feature space.
Original data
RBF Kernel PCA (γ=1)
Kernel matrix
200 points | Kernel: RBF exp(-γ‖x−y‖²), γ=1
from sklearn.decomposition import PCA, KernelPCA
from sklearn.datasets import make_circles
# Concentric circles: linear PCA fails, kernel PCA succeeds
X_circles, y_circles = make_circles(n_samples=500, factor=0.3, noise=0.05)
# Linear PCA cannot separate the two rings
Z_linear = PCA(n_components=2).fit_transform(X_circles)
# RBF Kernel PCA separates them
Z_rbf = KernelPCA(n_components=2, kernel='rbf', gamma=10).fit_transform(X_circles)
# The bandwidth γ controls locality:
# Small γ → global (nearly linear)
# Large γ → local (sharp boundaries)
for gamma in [0.1, 2, 10, 50]:
kpca = KernelPCA(n_components=2, kernel='rbf', gamma=gamma)
Z = kpca.fit_transform(X_circles)
print(f" γ = {gamma:5.1f}: separation quality varies")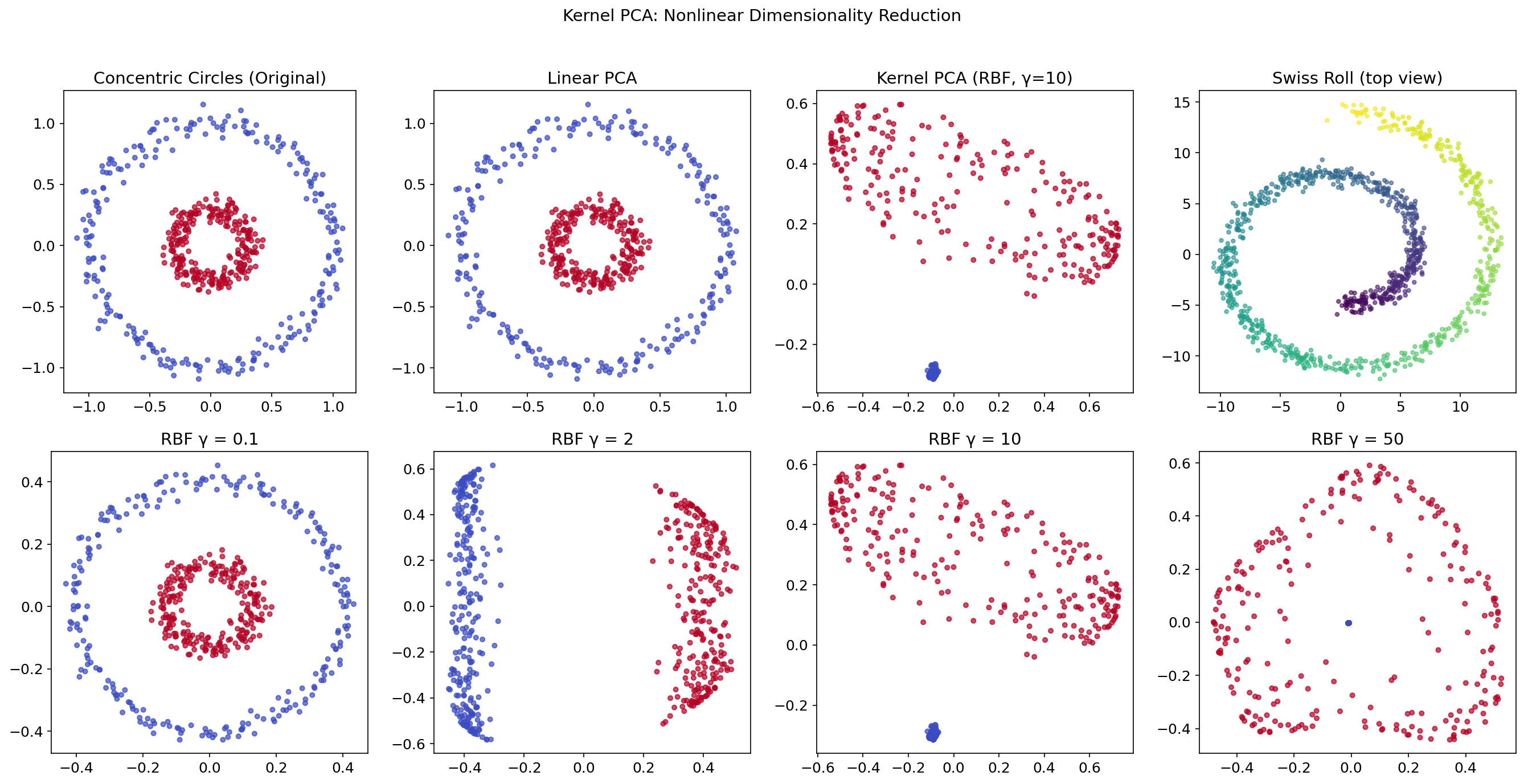
Probabilistic PCA
PCA as a Latent Variable Model
Probabilistic PCA (Tipping & Bishop, 1999) recasts PCA as a generative model:
where:
- is a -dimensional latent variable.
- is the loading matrix.
- is the data mean.
- is isotropic Gaussian noise.
Theorem 4 (MLE for Probabilistic PCA).
The maximum likelihood estimates are:
where contains the top eigenvectors of , , is an arbitrary rotation, and:
The noise variance is the average of the discarded eigenvalues — the “lost” variance is modeled as isotropic noise.
Why Probabilistic PCA?
- Missing data: the EM algorithm naturally handles incomplete observations.
- Model comparison: the marginal likelihood allows Bayesian model selection for .
- Generative model: we can sample new data points from .
- Mixture of PPCA: captures multimodal structure (cluster-specific PCA).
def ppca_em(X, k, n_iter=100, missing_mask=None):
"""Probabilistic PCA via EM. missing_mask[i,j]=True means X[i,j] is missing."""
n, d = X.shape
mu = np.nanmean(X, axis=0) if missing_mask is not None else X.mean(axis=0)
W = np.random.randn(d, k) * 0.1
sigma2 = 1.0
for iteration in range(n_iter):
# E-step: posterior p(z|x) = N(M^{-1} W^T (x - mu), sigma^2 M^{-1})
M = W.T @ W + sigma2 * np.eye(k)
M_inv = np.linalg.inv(M)
Ez = np.zeros((n, k))
Ezz = np.zeros((n, k, k))
for i in range(n):
if missing_mask is not None and np.any(missing_mask[i]):
obs = ~missing_mask[i]
W_obs = W[obs, :]
x_obs = X[i, obs] - mu[obs]
M_obs = W_obs.T @ W_obs + sigma2 * np.eye(k)
Ez[i] = np.linalg.solve(M_obs, W_obs.T @ x_obs)
Ezz[i] = sigma2 * np.linalg.inv(M_obs) + np.outer(Ez[i], Ez[i])
else:
xi = X[i] - mu
Ez[i] = M_inv @ W.T @ xi
Ezz[i] = sigma2 * M_inv + np.outer(Ez[i], Ez[i])
# M-step: update W and sigma^2
S1 = np.zeros((d, k))
S2 = np.zeros((k, k))
for i in range(n):
xi = X[i] - mu
if missing_mask is not None and np.any(missing_mask[i]):
obs = ~missing_mask[i]
S1[obs] += np.outer(xi[obs], Ez[i])
else:
S1 += np.outer(xi, Ez[i])
S2 += Ezz[i]
W = S1 @ np.linalg.inv(S2)
# ... (noise variance update continues)
return W, sigma2, mu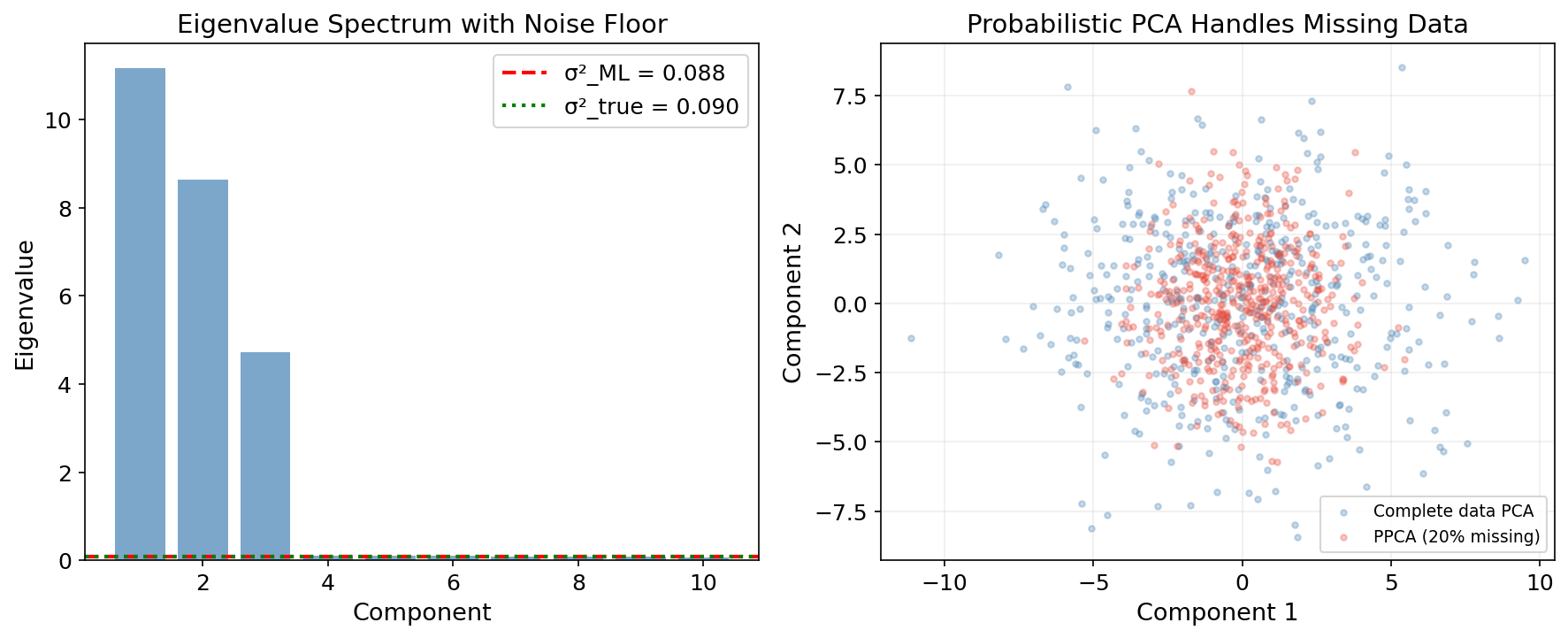
Robust PCA: Principal Component Pursuit
The Problem
Standard PCA is sensitive to gross corruption: a few large outliers or structured noise can distort the principal components. Robust PCA (Candès, Li, Ma & Wright, 2011) separates the data matrix into a low-rank component and a sparse component :
where captures the “clean” low-rank structure and captures sparse corruption (outliers, anomalies).
Principal Component Pursuit
Theorem 5 (Principal Component Pursuit).
Under mild incoherence conditions, the solution to:
recovers the true and exactly, where is the nuclear norm (sum of singular values) and is the entry-wise norm. The default parameter is .
The nuclear norm promotes low rank; the norm promotes sparsity. This convex relaxation can be solved efficiently by the Alternating Direction Method of Multipliers (ADMM).
Applications
- Video surveillance: = static background, = moving foreground objects.
- Face recognition: = faces under normal lighting, = shadows and occlusions.
- Financial data: = systematic market factors, = idiosyncratic shocks or anomalies.
def robust_pca_admm(X, lam=None, mu=None, tol=1e-7, max_iter=500):
"""Robust PCA via ADMM (Principal Component Pursuit).
Solves: min ||L||_* + λ||S||_1 s.t. L + S = X
"""
m, n = X.shape
if lam is None:
lam = 1.0 / np.sqrt(max(m, n))
if mu is None:
mu = m * n / (4.0 * np.sum(np.abs(X)))
L = np.zeros_like(X)
S = np.zeros_like(X)
Y = np.zeros_like(X) # dual variable
def soft_threshold(Z, threshold):
return np.sign(Z) * np.maximum(np.abs(Z) - threshold, 0)
def svd_threshold(Z, threshold):
U, sigma, Vt = np.linalg.svd(Z, full_matrices=False)
sigma_thresh = np.maximum(sigma - threshold, 0)
return U @ np.diag(sigma_thresh) @ Vt, np.sum(sigma_thresh > 0)
for iteration in range(max_iter):
# Update L (nuclear norm proximal operator)
L, rank_L = svd_threshold(X - S + Y / mu, 1.0 / mu)
# Update S (L1 proximal operator)
S = soft_threshold(X - L + Y / mu, lam / mu)
# Update dual variable
residual = X - L - S
Y = Y + mu * residual
if np.linalg.norm(residual, 'fro') / np.linalg.norm(X, 'fro') < tol:
break
return L, S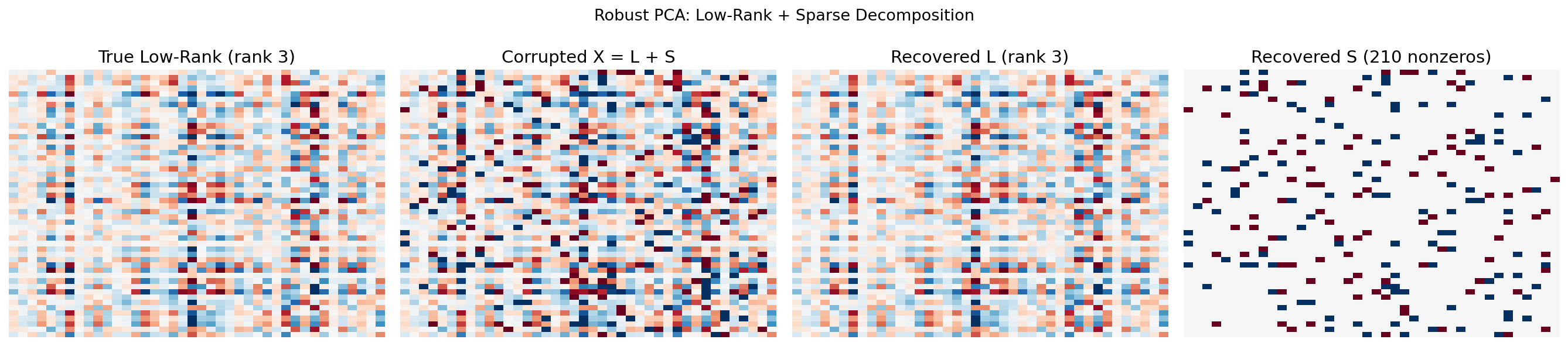
Sparse PCA
The Interpretability Problem
Standard PCA components are linear combinations of all features — every loading is typically nonzero. In high-dimensional settings (), this makes PCs hard to interpret: what does a component that loads on all 10,000 genes actually mean?
Sparse PCA Formulation
Sparse PCA adds an penalty to the loadings to encourage sparsity:
where controls the sparsity–variance trade-off. Larger → fewer nonzero loadings → more interpretable but less variance explained.
Definition 5 (Sparse PCA trade-off).
Sparse PCA sacrifices optimality (in variance explained) for interpretability (in sparsity of loadings). The trade-off is controlled by the regularization parameter :
- : standard PCA (dense loadings, maximum variance).
- : trivial solution (zero loadings).
- Intermediate : sparse loadings that explain most of the variance.
Several algorithms exist: SCoTLASS (Jolliffe et al., 2003), SPCA (Zou et al., 2006), and the coordinate descent approach in scikit-learn’s SparsePCA.
from sklearn.decomposition import SparsePCA, PCA
# Compare standard PCA vs. Sparse PCA on the Wine dataset
wine = load_wine()
X_std = StandardScaler().fit_transform(wine.data)
pca = PCA(n_components=3).fit(X_std)
spca = SparsePCA(n_components=3, alpha=1.0, random_state=42).fit(X_std)
# Standard PCA: all 13 features contribute to each component
print("Standard PCA loadings (PC 1):")
print(f" Nonzero: {np.sum(np.abs(pca.components_[0]) > 0.01)}/13")
# Sparse PCA: only a few features per component
print("Sparse PCA loadings (PC 1):")
print(f" Nonzero: {np.sum(np.abs(spca.components_[0]) > 0.01)}/13")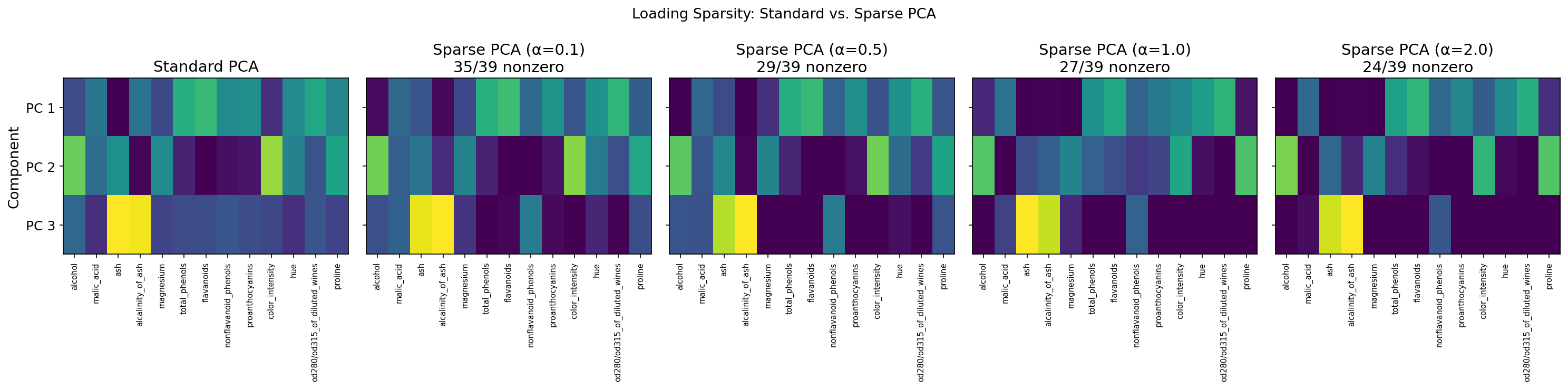
Connections & Further Reading
PCA stands at the crossroads of the Linear Algebra track and the rest of the formalML curriculum.
| Topic | PCA Connection |
|---|---|
| The Spectral Theorem | PCA eigendecomposition is the Spectral Theorem applied to . The Rayleigh quotient and Courant–Fischer theorem give the variance-maximization formulation. |
| Singular Value Decomposition | PCA is the SVD of the centered data matrix. The Eckart–Young theorem makes PCA the optimal low-rank projection. The SVD avoids squaring the condition number. |
| Tensor Decompositions | Tucker decomposition generalizes PCA to multi-way arrays. Multilinear PCA (MPCA) applies PCA along each tensor mode. |
| Persistent Homology | PCA is often a preprocessing step for TDA: reduce dimensionality before computing persistent homology. The explained variance ratio guides the choice of dimensionality. |
| The Mapper Algorithm | Mapper frequently uses PCA (via SVD) as a filter function — projecting data onto the first 1–2 principal components before the covering step. |
| Sheaf Theory | The Sheaf Laplacian’s eigendecomposition (Spectral Theorem) combined with PCA on the resulting feature vectors provides multi-scale structural analysis. |
| Measure-Theoretic Probability | The Law of Large Numbers justifies why ; the projection interpretation of conditional expectation is the infinite-dimensional analog of PCA’s variance-minimizing projection. |
The Linear Algebra Track Dependency Graph
Spectral Theorem
├── Singular Value Decomposition
│ └── PCA & Low-Rank Approximation (this topic)
└── Tensor DecompositionsPCA is the SVD applied to centered data. The Spectral Theorem provides the eigendecomposition of the covariance matrix. Tensor Decompositions generalize both to multilinear algebra.
Connections
- PCA is the SVD of the centered data matrix; the Eckart–Young theorem makes the truncated SVD the optimal low-rank projection; the SVD avoids squaring the condition number when computing PCA svd
- PCA eigendecomposition is the Spectral Theorem applied to the sample covariance matrix; the Rayleigh quotient and Courant–Fischer theorem give the variance-maximization formulation spectral-theorem
- Tucker decomposition generalizes PCA to multi-way arrays; Multilinear PCA (MPCA) applies PCA along each tensor mode tensor-decompositions
- PCA is a standard preprocessing step for TDA — reducing dimensionality before computing persistent homology persistent-homology
- Mapper frequently uses PCA (via SVD) as a filter function — projecting data onto the first 1–2 principal components before the covering step mapper-algorithm
- the Sheaf Laplacian eigendecomposition (Spectral Theorem) combined with PCA on the resulting feature vectors provides multi-scale structural analysis sheaf-theory
- the Law of Large Numbers guarantees that the sample covariance converges to the population covariance; the L2 projection interpretation of conditional expectation is the infinite-dimensional analog of PCA measure-theoretic-probability
- Projection depth (statistical-depth §3.3) is the worst-case standardized residual over all 1D projections — a robustified version of the leading PCA loading. PCA finds maximum-variance subspaces; projection depth finds maximum-outlyingness directions over the same sphere of unit projections. statistical-depth
- Calibration scores in conformal prediction function as low-dimensional summaries of prediction error; the leave-one-out updates underlying jackknife+ are formally analogous to the LOO covariance updates used in low-rank settings, with the same exchangeability + rank-symmetry machinery delivering finite-sample coverage. conformal-prediction
References & Further Reading
- book Introduction to Linear Algebra — Strang (2016) Chapter 7: SVD and PCA connection
- book Pattern Recognition and Machine Learning — Bishop (2006) Chapter 12: Probabilistic PCA and EM derivation
- book The Elements of Statistical Learning — Hastie, Tibshirani & Friedman (2009) Chapter 14.5: PCA, Sparse PCA, kernel PCA
- book Matrix Analysis — Horn & Johnson (2013) Rayleigh quotient, Courant–Fischer, and Eckart–Young proofs
- paper Probabilistic Principal Component Analysis — Tipping & Bishop (1999) Original PPCA paper
- paper Robust Principal Component Analysis? — Candès, Li, Ma & Wright (2011) Principal Component Pursuit
- paper Sparse Principal Component Analysis — Zou, Hastie & Tibshirani (2006) SPCA formulation and algorithm
- paper Nonlinear Component Analysis as a Kernel Eigenvalue Problem — Schölkopf, Smola & Müller (1998) Original Kernel PCA paper
- paper A generalization of principal component analysis to the exponential family — Collins, Dasgupta & Schapire (2001) Exponential family PCA
- paper A parallel analysis criterion for determining the number of common factors or principal components — Horn (1965) Parallel analysis for dimensionality selection